|
These versions allows owners of old Mac to continue to work with AmplifX: version 1.6.3 doesn’t force the user to instal the new 1.7.0 release. Manage, test and design your primers for PCR Vous cherchez la version française d’AmplifX : cliquer sur le lien "Français" en haut à droite de la page ! And a lot of bug fixes and algorithm corrections! The menu "paste complement strand" now works into the field "use this primer" in the primer design window. New interface element (slider) in the main window that allow to quickly change the required number of perfect matching 3’end nucleotides in the match search alogorithm (without going to the Preference window) Ability to change the primer order in the list by drag and drop Multi-criteria sorting of primer list (see the new menu "primers > Sort using multiple criteria.") The program is now distributed as a multilingual bundle (still only in english and french but translators are welcome) Huge improvment in speed for almost all time consuming operations: sequence and primer list loading, match searches, primer design. Very big sequences support (> 1 million nucleotides) The Mac version is now a Cocoa application (compatible Mac OS 10.9 Mavericks and Retina displays) 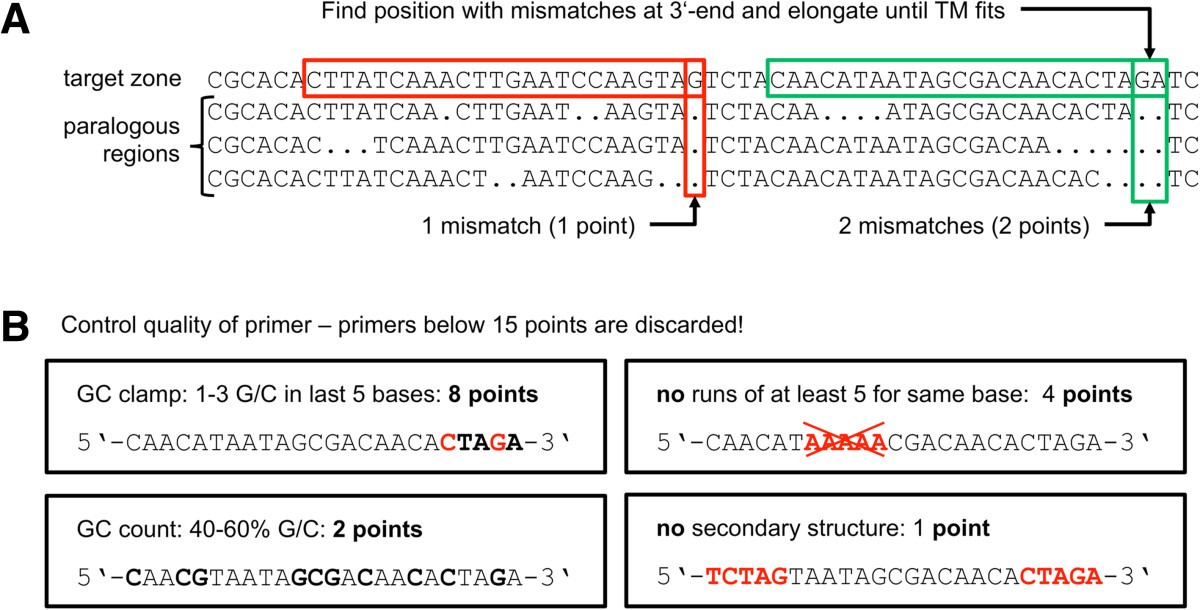
Simple and intuitive interface to design new primers for amplifying a fragment choosing the amplicon size and/or the position Pertinent informations on matches and amplicons (TM, size, predicted dimers, etc.) Now to design the primers using the 5-3 (sense) strand: One of the primers (the forward primer) will be directed from 5-3 and anneal with the antisense.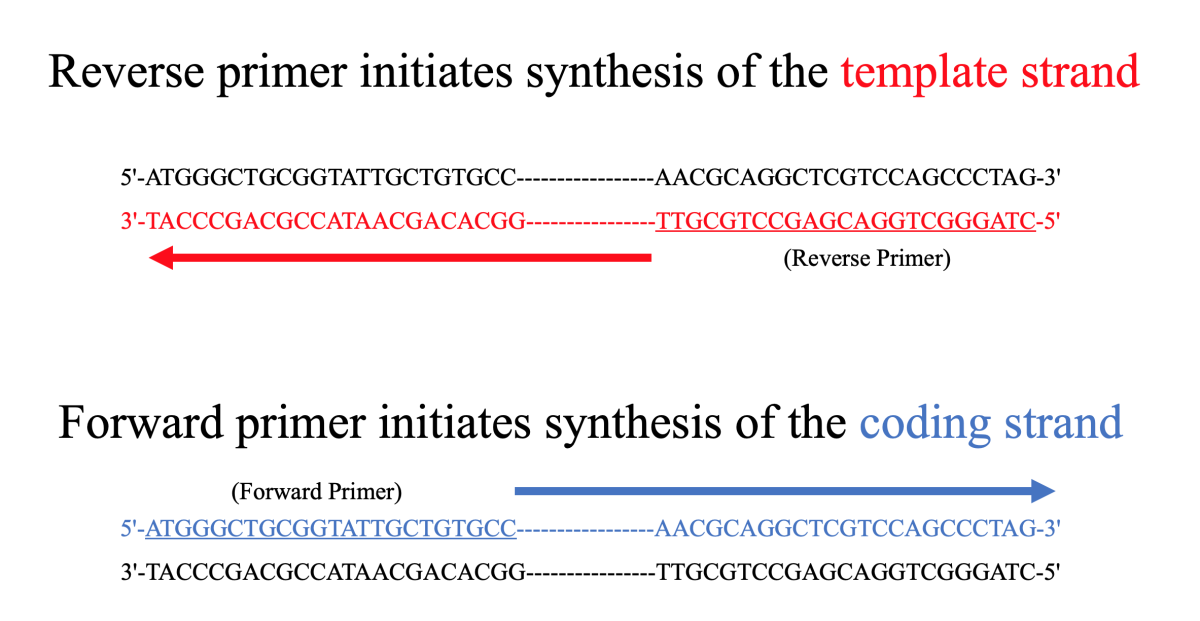  
Localise your primers on target sequences (main file formats supported and "intelligent" copy-paste) Share simply primer lists (automatic lockout in read only mode of files yet opened by an other user)Īutomatic calculation of quality score (TM, length, GC%, autodimer, complexity, etc.)
0 Comments
Leave a Reply. |
Details
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |

 RSS Feed
RSS Feed
